-Search query
-Search result
Showing 1 - 50 of 234 items for (author: de & simone & a)
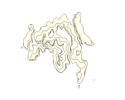
EMDB-19184: 
Late alpha-Synuclein fibril structure from liquid-liquid phase separations.
Method: helical / : De Simone A, Barritt JD, Chen S, Cascella R, Cecchi C, Bigi A, Jarvis JA, Chiti F, Dobson CM, Fusco G

PDB-8ri9: 
Late alpha-Synuclein fibril structure from liquid-liquid phase separations.
Method: helical / : De Simone A, Barritt JD, Chen S, Cascella R, Cecchi C, Bigi A, Jarvis JA, Chiti F, Dobson CM, Fusco G

EMDB-40603: 
GPR161 Gs heterotrimer
Method: single particle / : Hoppe N, Manglik A, Harrison S

PDB-8smv: 
GPR161 Gs heterotrimer
Method: single particle / : Hoppe N, Manglik A, Harrison S

EMDB-16229: 
Cryo-EM structure of the bacterial replication origin opening basal unwinding system
Method: single particle / : Pelliciari S, Bodet-Lefevre S, Murray H, Ilangovan A

PDB-8btg: 
Cryo-EM structure of the bacterial replication origin opening basal unwinding system
Method: single particle / : Pelliciari S, Bodet-Lefevre S, Murray H, Ilangovan A

EMDB-16328: 
Outer membrane attachment porin OmpM1 from Veillonella parvula
Method: single particle / : Silale A, van den Berg B

EMDB-16332: 
Outer membrane attachment porin OmpM1 from Veillonella parvula, native
Method: single particle / : Silale A, van den Berg B

EMDB-16333: 
Outer membrane attachment porin OmpM1 from Veillonella parvula, C3 symmetry
Method: single particle / : Silale A, van den Berg B

PDB-8bym: 
Outer membrane attachment porin OmpM1 from Veillonella parvula
Method: single particle / : Silale A, van den Berg B

PDB-8bys: 
Outer membrane attachment porin OmpM1 from Veillonella parvula, native
Method: single particle / : Silale A, van den Berg B

PDB-8byt: 
Outer membrane attachment porin OmpM1 from Veillonella parvula, C3 symmetry
Method: single particle / : Silale A, van den Berg B

EMDB-16890: 
Iron Nitrogenase Complex from Rhodobacter capsulatus
Method: single particle / : Schmidt FV, Schulz L, Zarzycki J, Prinz S, Erb TJ, Rebelein JG

EMDB-17583: 
CHAPSO treated partial catalytic component (comprising only AnfD & AnfK, lacking AnfG and FeFeco) of iron nitrogenase from Rhodobacter capsulatus
Method: single particle / : Schmidt FV, Schulz L, Zarzycki J, Prinz S, Erb TJ, Rebelein JG

PDB-8oie: 
Iron Nitrogenase Complex from Rhodobacter capsulatus
Method: single particle / : Schmidt FV, Schulz L, Zarzycki J, Prinz S, Erb TJ, Rebelein JG

PDB-8pbb: 
CHAPSO treated partial catalytic component (comprising only AnfD & AnfK, lacking AnfG and FeFeco) of iron nitrogenase from Rhodobacter capsulatus
Method: single particle / : Schmidt FV, Schulz L, Zarzycki J, Prinz S, Erb TJ, Rebelein JG
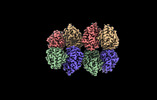
EMDB-28013: 
Cryo-EM structure of the Glutaminase C core filament (fGAC)
Method: single particle / : Ambrosio AL, Dias SM, Quesnay JE, Portugal RV, Cassago A, van Heel MG, Islam Z, Rodrigues CT

PDB-8ec6: 
Cryo-EM structure of the Glutaminase C core filament (fGAC)
Method: single particle / : Ambrosio AL, Dias SM, Quesnay JE, Portugal RV, Cassago A, van Heel MG, Islam Z, Rodrigues CT

EMDB-40789: 
BAP1/ASXL1 bound to the H2AK119Ub Nucleosome
Method: single particle / : Thomas JF, Valencia-Sanchez MI, Armache KJ

EMDB-40790: 
Map focused on acidic patch BAP1/ASXL1 bound to the H2AK119Ub Nucleosome
Method: single particle / : Thomas JF, Valencia-Sanchez MI

EMDB-40791: 
Overall map of BAP1/ASXL1 bound to the H2AK119Ub Nucleosome
Method: single particle / : Thomas JF, Valencia-Sanchez MI

PDB-8svf: 
BAP1/ASXL1 bound to the H2AK119Ub Nucleosome
Method: single particle / : Thomas JF, Valencia-Sanchez MI, Armache KJ

EMDB-17756: 
Structure of the murine trace amine-associated receptor TAAR7f bound to N,N-dimethylcyclohexylamine (DMCH) in complex with mini-Gs trimeric G protein
Method: single particle / : Gusach A, Lee Y, Edwards PC, Huang F, Weyand SN, Tate CG

PDB-8pm2: 
Structure of the murine trace amine-associated receptor TAAR7f bound to N,N-dimethylcyclohexylamine (DMCH) in complex with mini-Gs trimeric G protein
Method: single particle / : Gusach A, Lee Y, Edwards PC, Huang F, Weyand SN, Tate CG

EMDB-27121: 
CryoEM structure of human orphan GPCR GPR179 in complex with extracellular matrix protein pikachurin
Method: single particle / : Patil DN, Martemyanov KA

PDB-8d1b: 
CryoEM structure of human orphan GPCR GPR179 in complex with extracellular matrix protein pikachurin
Method: single particle / : Patil DN, Martemyanov KA

EMDB-40184: 
Structure of the Spizellomyces punctatus Fanzor (SpuFz) in complex with omega RNA and target DNA
Method: single particle / : Xu P, Saito M, Zhang F

PDB-8gkh: 
Structure of the Spizellomyces punctatus Fanzor (SpuFz) in complex with omega RNA and target DNA
Method: single particle / : Xu P, Saito M, Zhang F

EMDB-17540: 
Cryo-EM structure of full-length human UBR5 (homotetramer)
Method: single particle / : Aguirre JD, Kater L, Kempf G, Cavadini S, Thoma NH

PDB-8p83: 
Cryo-EM structure of full-length human UBR5 (homotetramer)
Method: single particle / : Aguirre JD, Kater L, Kempf G, Cavadini S, Thoma NH

EMDB-17542: 
Negative stain map of UBR5 (dimer) in complex with RARA/RXRA
Method: single particle / : Aguirre JD, Cavadini S, Kempf G, Kater L, Thoma NH

EMDB-16376: 
CryoEM structure of a tungsten-containing aldehyde oxidoreductase from Aromatoleum aromaticum
Method: single particle / : Winiarska A, Ramirez-Amador F, Hege D, Gemmecker Y, Prinz S, Hochberg G, Heider J, Szaleniec M, Schuller JM

PDB-8c0z: 
CryoEM structure of a tungsten-containing aldehyde oxidoreductase from Aromatoleum aromaticum
Method: single particle / : Winiarska A, Ramirez-Amador F, Hege D, Gemmecker Y, Prinz S, Hochberg G, Heider J, Szaleniec M, Schuller JM

EMDB-17154: 
Cryo-EM structure of CLOCK-BMAL1 bound to a nucleosomal E-box at position SHL+5.8 (consensus and constituent map 1)
Method: single particle / : Stoos L, Michael AK, Kempf G, Cavadini S, Thoma NH

EMDB-17155: 
Cryo-EM structure of CLOCK-BMAL1 bound to a nucleosomal E-box at position SHL-6.2 (DNA conformation 1)
Method: single particle / : Michael AK, Stoos L, Kempf G, Cavadini S, Thoma NH

EMDB-17156: 
Cryo-EM structure of CLOCK-BMAL1 bound to a nucleosomal E-box at position SHL-6.2 (DNA conformation 2)
Method: single particle / : Michael AK, Stoos L, Kempf G, Cavadini S, Thoma NH

EMDB-17157: 
Cryo-EM structure of CLOCK-BMAL1 bound to a nucleosomal E-box at position SHL+5.8 (composite map)
Method: single particle / : Stoos L, Michael AK, Kempf G, Cavadini S, Thoma NH

EMDB-17158: 
Cryo-EM structure of CLOCK-BMAL1 bound to a nucleosomal E-box at position SHL+5.8 (constituent map 2 from additional focus classification on PAS domains)
Method: single particle / : Stoos L, Michael AK, Kempf G, Cavadini S, Thoma NH

EMDB-17159: 
Cryo-EM map of MYC-MAX-OCT4-LIN28 complex
Method: single particle / : Michael AK, Kempf G, Cavadini S, Thoma NH

EMDB-17160: 
Cryo-EM structure of CLOCK-BMAL1 bound to the native Por enhancer nucleosome (map 2, additional 3D classification and flexible refinement)
Method: single particle / : Michael AK, Stoos L, Kempf G, Cavadini S, Thoma N

EMDB-17161: 
Cryo-EM structure of CLOCK-BMAL1 bound to the native Por enhancer nucleosome (map 1)
Method: single particle / : Michael AK, Stoos L, Cavadini S, Kempf G

EMDB-17162: 
MAX-MAX bound to a nucleosome at SHL+5.1 and SHL-6.9.
Method: single particle / : Stoos L, Kempf G, Kater L, Thoma NH

EMDB-17183: 
OCT4 and MYC-MAX co-bound to a nucleosome
Method: single particle / : Michael AK, Stoos L, Kempf G, Cavadini S, Thoma N

EMDB-17184: 
MYC-MAX bound to a nucleosome at SHL+5.8
Method: single particle / : Stoos L, Michael AK, Kempf G, Kater L, Cavadini S, Thoma N

PDB-8osj: 
Cryo-EM structure of CLOCK-BMAL1 bound to a nucleosomal E-box at position SHL-6.2 (DNA conformation 1)
Method: single particle / : Michael AK, Stoos L, Kempf G, Cavadini S, Thoma NH

PDB-8osk: 
Cryo-EM structure of CLOCK-BMAL1 bound to a nucleosomal E-box at position SHL+5.8 (composite map)
Method: single particle / : Stoos L, Michael AK, Kempf G, Cavadini S, Thoma NH

PDB-8osl: 
Cryo-EM structure of CLOCK-BMAL1 bound to the native Por enhancer nucleosome (map 2, additional 3D classification and flexible refinement)
Method: single particle / : Michael AK, Stoos L, Kempf G, Cavadini S, Thoma N

PDB-8ots: 
OCT4 and MYC-MAX co-bound to a nucleosome
Method: single particle / : Michael AK, Stoos L, Kempf G, Cavadini S, Thoma N

PDB-8ott: 
MYC-MAX bound to a nucleosome at SHL+5.8
Method: single particle / : Stoos L, Michael AK, Kempf G, Kater L, Cavadini S, Thoma N

EMDB-15295: 
African cichlid nackednavirus capsid at pH 7.5
Method: single particle / : Pfister S, Rabl J, Boehringer D, Meier BH
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
